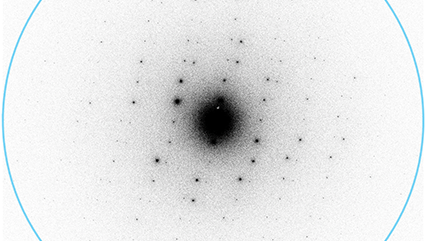
K3® 是直接探测相机性能表现的新标杆,从头开始重新设计,这款真正的新一代相机面向生命科学和材料科学研究领域极具挑战的低剂量电子显微术应用进行了优化。 K3 相机是Gatan 在实时、单电子计数直接探测相机技术领域深厚经验的完整与全新表达。
- 强大的在线信号处理功能,带来超越 K2® 相机的 DQE 表现; CDS 模式的加入进一步提高相机的DQE
- 实时电子计数使您可以即时判断样品质量
- 可选配基于 GPU 的在线运动修正模块,避免储存体量高达 TB 级的原始图像帧数据
- 每秒 1500 帧全幅读出 – K2 相机的 3.75 倍
- 匹配您应用需要的视野大小
- 2400 万像素— – 为您的高性能电镜提供更高的成像通量
- 1400 万像素— – 将您的筛样电镜变为数据采集电镜
-
 K3 camera for MicroED/3DED
K3 camera for MicroED/3DED
-
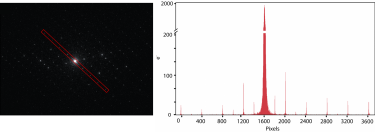 Electron-counted SAED of ZSM-5 with the K3 camera
Electron-counted SAED of ZSM-5 with the K3 camera
-
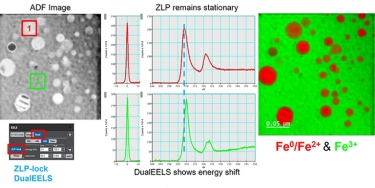
 DualEELS:对低能损电子能量损失谱数据校正的重要性
DualEELS:对低能损电子能量损失谱数据校正的重要性
-
 Imaging molecules in their native environment: Cryo-electron tomography of PCDH15 complexes in mouse stereocilia
Imaging molecules in their native environment: Cryo-electron tomography of PCDH15 complexes in mouse stereocilia
-
 Cryo EM reveals mechanisms of gating and drug modulation in 5 HT3A receptors webinar
Cryo EM reveals mechanisms of gating and drug modulation in 5 HT3A receptors webinar
-
 CryoARM/K3 组合带来的高解析分辨率:SerialEM,Latitude以及数据采集未来发展方向
CryoARM/K3 组合带来的高解析分辨率:SerialEM,Latitude以及数据采集未来发展方向
-

 结构生物学助力疫情应对
结构生物学助力疫情应对
-

 NUANCE Workshop on 4D STEM: Fundamentals of Electron Diffraction and 4D STEM
NUANCE Workshop on 4D STEM: Fundamentals of Electron Diffraction and 4D STEM
-

 NUANCE Workshop on 4D STEM: Data Processing in DM
NUANCE Workshop on 4D STEM: Data Processing in DM
-

 NUANCE Workshop on 4D STEM: Data Processing using Python
NUANCE Workshop on 4D STEM: Data Processing using Python
In situ structural analysis of SARS-CoV-2 spike reveals flexibility mediated by three hinges
Turoňová, B.; Sikora, M.; Schürmann, C.; Hagen, W. J. H.; Welsch, S.; Blanc, F. E. C.; von Bülow, S.; Gecht, M.; Bagola, K.; Hörner, C.; van Zandbergen, G.; Landry, J.; de Azevedo, N. T. D.; Mosalaganti, S.; Schwarz, A.; Covino, R.; Mühlebach, M. D.; Hummer, G.; Locker, J. K.; Beck, M.
Molecular architecture of the SARS-CoV-2 virus
Yao, H.; Song, Y.; Chen, Y.; Wu, N.; Xu, Sun, C.; Zhang, J.; Weng, T.; Zhang, Z.; Wu, Z.; Cheng, L,; Shi, D.; Lu, X.; Lei, J.; Crispin, M.; Shi, Y.; Li, L.; Li, S.
Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation
Wrapp, D.; Wang, N.; Corbett, K. S.; Goldsmith, J. A.; Hsieh, C. -L.; Abiona, O.; Graham, B. S.; McLellan, J. S.
Time-resolved cryo-EM using a combination of droplet microfluidics with on-demand jetting
Torino, S.; Dhurandhar, M.; Stroobants, A.; Claessens, R.; Efremov, R. G.
Szemán, A. J. K.; Stráner, P.; Jákli, I.; Hosogi, N.; Harmat, V.; Menyhárd, D. K.; Perczel, A.
Structural basis of actin filament assembly and aging
Oosterheert, W.; Klink, B. U.; Belyy, A.; Pospich, S.; Raunser, S.
Helical ultrastructure of the metalloprotease meprin α in complex with a small molecule inhibitor
Bayly-Jones, C.; Lupton, C. J.; Fritz, C.; Venugopal, H.; Ramsbeck, D.; Wermann, M.; Jäger,C.; Marco, A. D.; Schilling, S.; Schlenzig, D.; Whisstock, J. C.
Stern, A. M.; Yang, Y.; Meunier, A. L.; Liu, W.; Cai, Y.; Ericsson, M.; Liu, L.; Goedert, M.; Scheres, S. H. W.; Selkoe, D.J
Cryo-EM structure of the human NKCC1 transporter reveals mechanisms of ion coupling and specificity
Neumann, C.; Rosenbæk, L. L.; Flygaard, R. K.; Habeck, M.; Karlsen, J. L.; Wang, Y.; Larsen, K. L.; Gad, H. H.; Hartmann, R.; Lyons, J. A; Fenton, R. A.; Nissen, P.
Algal photosystem I dimer and high-resolution model of PSI-plastocyanin complex
Naschberger, A.; Mosebach, L.; Tobiasson, V.; Kuhlgert, S.; Scholz, M.; Perez-Boerema, A.; Ho, T. T. H.; Vidal-Meireles, A.; Takahashi, Y.; Hippler, M.; Amunts, A.
Chang, M. R.; Tomasovic, L.; Kuzmina, N. A.; Ronk, A. J.; Byrne, P. O.; Johnson, R.; Storm, N.; Olmedillas, E.; Hou, Y. J.; Schäfer, A.; Leist, S. R.; Tse, L. T.; Ke, H.; Coherd, C.; Nguyen, K.; Kamkaew, M.; Honko, A.; Zhu, Q.; Alter, G.; Saphire, E. O.; McLellan, J. S.; Griffiths, A.; Baric, R. S.; Bukreyev. A.; Marasco. W. A.
Frieg, B.; Geraets, J. A.; Strohäker, T.; Dienemann, C.; Mavroeidi, P.; Jung, B. C.; Kim, W. S.; Lee, S. J.; Xilouri, M.; Zweckstetter. M.; Schröder. G. F.
Organic crystal growth: Hierarchical self-assembly involving nonclassical and classical steps
Biran, I.; Rosenne, S.; Weissman, H.; Tsarfati, Y.; Houben, L.; Rybtchinski. B.
Lyu, M.; Ayala, J. C.; Chirakos, I.; Su, C. -C.; Shafer, W. M.; Yu, E. W.
Structure of the PAPP-ABP5 complex reveals mechanism of substrate recognition
Judge, R. A.; Sridar, J.; Tunyasunvunakool, K.; Jain, R.; Wang, J. C. K.; Ouch, C.; Xu, J.; Mafi, A.; Nile, A. H.; Remarcik, C.; Smith, C. L.; Ghosh, C.; Xu, C.; Stoll, V.; Jumper, J.; Singh, A. H.; Eaton, D.; Hao, Q.
Deagglomeration of DNA nanomedicine carriers using controlled ultrasonication
Hinchliffe, B. A.; Turner, P.; J. H. Cant, D.; De Santis, E.; Aggarwal, P.; Harris, R.; Templeton, D.; Shard, A. G.; Hodnett, M.; Minelli, C.
Structures of a phycobilisome in light-harvesting and photoprotected states
Domínguez-Martín, M. A.; Sauer, P. V.; Kirst, H.; Sutter, M.; Bína, D.; Greber, B. J.; Nogales, E. ;Polívka, T.; Kerfeld, C. A.
Zhu, J.; Huang, W.; Zhao, J., Huynh, L.; Taylor, D. J.; Harris, M. E.
Miyakawa, T.; Yang, J.; Kawasaki, M.; Adachi, N.; Fujii, A.; Miyauchi, Y.; Muramatsu, T.; Moriya, T.; Senda, T.; Tanokura, M.
Zhao, J.; Makhija, S.; Zhou, C.; Zhang, H.; Wang, Y.; Muralidharan, M.; Huang, B.; Cheng, Y.
Mechanistic details of CRISPR-associated transposon recruitment and integration revealed by cryo-EM
Park, J. U.; Tsai, A. W. -T.; Chen, T. H.; Peters, J. E.; Kellogg, E. H.
Specific recognition and ubiquitination of slow-moving ribosomes by human CCR4-NOT
Absmeier, E.; Chandrasekaran, V.; O'Reilly, F. J.; Stowell, J. A. W.; Rappsilber, J.; Passmore, L. A.
Cryo-electron microscopy of extracellular vesicles
Cai, K.; Sibert, B. S.; Kumar, A.; Yang, J.; Larson, M.; Thompson, K.; Wright, E. R.
Peck, J. V.; Strauss, J. D.; Fay, J. F.
Cryo-EM structure of an active bacterial TIR–STING filament complex
Morehouse, B. R.; Yip, M. C. J.; Keszei, A. F. A.; McNamara-Bordewick, N. K.; Shao, S.; Kranzusch, P. J.
Maintaining the momentum in cryoEM for biological discovery
Thompson, R.; Halfon, Y.; Aspinall, L.; White, J.; Hirst, I. J.; Wang, Y.; Darrow, M.; Muench, S. P.
Structural analysis of the basal state of the Artemis:DNA-PKcs complex
Watanabe, G.; Lieber, M. R.; Williams, D. R.
Flipped over U: structural basis for dsRNA cleavage by the SARS-CoV-2 endoribonuclease
Frazier, M. N.; Wilson, I. M.; Krahn, J. M.; Butay, K. J.; Dillard, L. B.; Borgnia, M. J.; Stanley, R. E.
Electron-counting MicroED data with the K2 and K3 direct electron detectors
Clabbers, M. T. B.; Martynowycz, M. W.; Hattne, J.; Nannenga, B. L.; Gonen, T.
Structures and gating mechanisms of human bestrophin anion channels
Owji, A. P.; Wang, J.; Kittredge, A.; Clark, Z.; Zhang, Y.; Hendrickson, W. A.; Yang, T.
Structural insights into dsRNA processing by Drosophila Dicer-2–Loqs-PD
Su, S.; Wang, J.; Deng, T.; Yuan, X.; He, J.; Liu, N.; Li, X.; Huang, Y.; Wang, H. -W.; Ma, J.
Structures and mechanism of the plant PIN-FORMED auxin transporter
Ung, K. L.; Winkler, M.; Schulz, L.; Kolb, M.; Janacek, D. P.; Dedic, E.; Stokes, D. L.; Hammes, U. Z.; Pedersen, B. P.
Structure and flexibility of the yeast NuA4 histone acetyltransferase complex
Zukin, S. A.; Marunde, M. R.; Popova, I. K.; Nogales, E.; Patel, A. B.
Role of aIF5B in archaeal translation initiation
Kazan, R.; Bourgeois, G.; Lazennec-Schurdevin, C.; Larquet, E.; Mechulam, Y.; Coureux, P. -D.; Schmitt, E.
Ion complexation waves emerge at the curved interfaces of layered minerals
Whittaker, M. L.; Ren, D.; Ophus, C.; Zhang, Y.; Waller, L.; Gilbert, B.; Banfield, J. F.
Compact IF2 allows initiator tRNA accommodation into the P site and gates the ribosome to elongation
Basu, R. S.; Sherman, M. B.; Gagnon, M. G.
Structural Basis for pH-gating of the K+ channel TWIK1 at the selectivity filter
Turney, T. S.; Li, V.; Brohawn, S. G.
Cryo-EM structures of SARS-CoV-2 Omicron BA.2 spike
Stalls, V.; Lindenberger, J.; Gobeil, S. M. -C.; Henderson, R.; Parks, R.; Barr, M.; Deyton, M.; Martin, M.; Janowska, K.; Huang, X.; May, A.;l Speakman, M.; Beaudoin, E.; Kraft, B.; Lu, X.; Edwards, R. J.; Eaton, A.; Montefiori, D. C.; Williams, W.; Saunders, K. O.; Wiehe, K.; Haynes, B. F.; Acharya, P.
Structure of S1PR2–heterotrimeric G13 signaling complex
Chen, H.; Chen, K.; Huang, W.; Staudt, L. M.; Cyster, J. G.; Li, X.
Ishii, N.
Structure of the type V-C CRISPR-Cas effector enzyme
Kurihara, N.; Nakagawa, R.; Hirano, H.; Okazaki, S.; Tomita, A.; Kobayashi, K.; Kusakizako, T.; Nishizawa, T.; Yamashita, K.; Scott, D. A.; Nishimasu, H.; Nureki, O.
Zhao, W.; Jensen, G. J.
The giant Mimivirus 1.2 Mb genome is elegantly organized into a 30 nm helical protein shield
Villalta, A.; Schmitt, A.; Estrozi, L. F.; Quemin, E. R. J.; Alempic, J. -M.; Lartigue, A.; Pražák, V.; Belmudes, L.; Vasishtan, D.; Colmant, A. M. G.; Honoré, F. A.; Couté, Y.; Grünewald, K.; Abergel, C.
Structural and functional impact by SARS-CoV-2 Omicron spike mutations
Zhang, J.; Cai, Y.; Lavine, C. L.; Peng, H.; Zhu, H.; Anand, K.; Tong, P.; Gautam, A.; Mayer, M. L.; Rits-Volloch, S.; Wang, S.; Sliz, P.; Wesemann, D. R.; Yang, W.; Seaman, M. S.; Lu, J.; Xiao, T.; Chen, B.
Yu, H.; Hamaguchi, T.; Nakajima, Y.; Kato, K.; Kawakami, K.; Akita, F.; Yonekura, K.; Shen, J. -R.
Ultrapotent antibodies against diverse and highly transmissible SARS-CoV-2 variants
Wang, L.; Zhou, T.; Zhang, Y.; Yang, E. S.; Schramm, C. A.
Native-like SARS-CoV-2 spike glycoprotein expressed by ChAdOx1 nCoV-19/AZD1222 vaccine
Watanabe, Y.; Mendonça, L.; Allen, E. R.; Howe, A.; Lee, M.; Allen, J. D.; Chawla, H.; Pulido, D.; Donnellan, F.; Davies, H.; Ulaszewska, M.; Belij-Rammerstorfer, S.; Morris, S.; Krebs, A. -S.; Dejnirattisai, W.; Mongkolsapaya, J.; Supasa, P.; Screaton, G. R.; Green, C. M.; Lambe, T.; Zhang, P.; Gilbert, S. C.; Crispin, M.
Structure of a microtubule-bound axonemal dynein
Walton, T.; Wu, H.; Brown, A.
Structure-guided multivalent nanobodies block SARS-CoV-2 infection and suppress mutational escape
Koenig, P. -D.; Das, H.; Liu H.; Kümmerer, B. M.; Gohr, F. N.; Jenster, L. -M.; Schiffelers, L. D. J.; Tesfamariam, Y. M.; Uchima, M.; Wuerth, J. D.; Gatterdam, K.; Ruetalo, N.; Christensen, M. H.; Fandrey, C. I.; Normann, S.; Tödtmann, J.; M. P.; Pritzl, S.; Hanke, L.; Boos, J.; Yuan, M.; Zhu, X.; Schmid-Burgk, J. L.; Kato, H.; Schindler, M.; Wilson, I. A.; Geyer, M.; Ludwig, K. U.; Hällberg, M.; Wu, N. C.; Schmidt, F. I.
Mechanism of SARS-CoV-2 polymerase stalling by remdesivir
Kokic, G.; Hillen, H. S.; Tegunov, D.; Dienemann, C.; Seitz, F.; Schmitzova, J.; Farnung, L.; Siewert, A.; Höbartner, C.; Cramer, P.
Stabilizing the closed SARS-CoV-2 spike trimer
Juraszek, J.; Rutten, L.; Blokland, S.; Bouchier, P.; Voorzaat, R.; Ritschel, T.; Bakkers, M. J. G.; Renault , L. L. R.; Langedijk, J. P. M.
Zhou, T.; Tsybovsky, Y.; Gorman, J.; Rapp, M.; Cerutti, G.; Chuang, G. -Y.; Katsamba, P. S.; Sampson, J. M.; Schön, A.; Bimela, J.; Boyington, J. C.; Nazzari, A.; Olia, A. S.; Shi, W.; Sastry, M.; Stephens, T.; Stuckey, J.; Teng, I. -T.; Kwong, P. D
Architecture of a SARS-CoV-2 mini replication and transcription complex
Yan, L.; Zhang, Y.; Ge, J.; Zheng, L.; Gao, Y.; Wang, T.; Jia, Z.; Wang, H.; Huang, Y.; Li, M.; Wang, Q.; Ra, Z.; Lou, Z.
An ultrapotent synthetic nanobody neutralizes SARS-CoV-2 by stabilizing inactive spike
Schoof, M.; Faust, B.; Saunders, R. A.; Sangwan, S.; Rezelj, V.; Hoppe, N.; Boone, M.; Billesbølle, C. B.; Puchades, C.; Azumaya, C. M.; Kratochvil, H. T.; Zimanyi, M.; Deshpande, I.; Liang, J.; Dickinson, S.; Nguyen, H. C.; Chio, C. M.; Merz, G. E.; Thompson, M. C.; Diwanji, D.; Schaefer, K.; Anand, A. A.; Dobzinski, N.; Zha, B. S.; Simoneau, C. R.; Leon, K.; White, K. M.; Chio, U. S.; Gupta, M.; Jin, M.; Li, F.; Liu, Y.; Zhang, K.; Bulkley, D.; Sun, M.; Smith, A. M.; Rizo, A. N.; Moss, F.; Brilot, A. F.; Pourmal, S.; Trenker, R.; Pospiech, T.; Gupta, S.; Barsi-Rhyne, B.; Belyy, V.; Barile-Hill, A. W.; Nock, S.; Liu, Y.; Krogan, N. J.; Ralston, C. Y.; Swaney, D. L.; García-Sastre, A.; Ott, M.; Vignuzzi, M.; QCRG Structural Biology Consortium; Walter, P.; Manglik, A.
Zhou, T.; Teng, I. -T.; Olia, A. S.; Cerutti, G.; Gorman, J.; Nazzari, A.; Shi, W.; Tsybovsky, Y.; Wang, L.; Wang, S.; Zhang, B.; Zhang, Y.; Katsamba, P. S.; Petrova, Y.; Banach, B. B.; Fahad, A. S.; Liu, L.; Lopez Acevedo, S. N.; Madan, B.; de Souza, M. O.; Pan, X.; Wang, P.; Wolfe, J. R.; Yin, M.; Ho, D. D.; Phung, E.; DiPiazza, A.; Chang, L. A.; Abiona, O. M.; Corbett, K. S.; DeKosky, B. J.; Graham, B. S.; Mascola, J. R.; Misasi, J.; Ruckwardt, T.; Sullivan, N. J.; Shapiro, L.; Kwong, P. D.
Langer, L. M.; Gat, Y.; Bonneau, F.; Conti, E.
Structural basis for the recognition of the SARS-CoV-2 by full-length human ACE2
Yan, R.; Zhang, Y.; Li, Y.; Xia, L.; Guo, Y.; Zhou, Q.
Surpassing the physical Nyquist limit to produce super-resolution cryo-EM reconstructions
Feathers, J. R.; Spoth, K. A.; Fromme, J. C.
1025 型
产品说明书
应用
相关产品
BioContinuum® 能量过滤器
Alpine® 直接探测相机
K3 IS 相机
Latitude® S 软件
Elsa™冷冻传输样品杆
相关材料
Nyquist 频率
剂量分割和颗粒运动校正
使用计数和超分辨率模式提高 DQ
| Modulation transfer function (MTF) curves | ||
|---|---|---|
| 200 kV | 300 kV | |
|
K3 |
CDS | CDS |
| Standard | Standard | |
致谢
K3 延续了我们成功合作打造 K2 的辉煌成果,是劳伦斯伯克利国家实验室 Peter Denes 团队和 David Agard 共同打造的成功成果。

