Observing beam-induced dendritic growth over two different timescales
Introduction
In-situ observations of dynamic processes in the electron microscope have been made for many decades and began shortly after the development of the first electron microscopes. The instrumentation available to record these dynamics has improved dramatically over the past several decades so that now it is routine to capture videos at frame rates faster than the human eye can process. The latest Gatan camera for in-situ transmission electron microscopy (TEM) not only extends this trend of increased speed but also enables high-speed data capture with as few electrons as possible via direct detection, real-time electron counting.
In this application note, we present low-dose observations of beam-induced dendritic growth. Cu metal dendrites grow as a metal-organic precursor is reduced by the electron beam. Two distinct experiments were performed. In the first experiment, the camera was used to capture rapid dynamics at 75 frames per second (fps). In the second, the same camera was used to capture much slower growth at a lower dose and frame rates. Analysis of growth rates from the first dataset is presented, indicating the possibilities enabled by the recent integration of Python scripting into the DigitalMicrograph® software.
High-speed observation
For the first experiment presented here, the goal was to capture dynamics at high speed, with minimum electron dose. To obtain this result, the magnification, dose rates at both sample and camera, and the frame rate need to be set correctly. The TEM magnification should be set as low as possible without losing the features of interest since low magnification allows the dose rate per pixel on the camera to be high relative to the number of electrons per square angstrom on the sample. This condition minimizes electron beam effects on the sample while simultaneously maximizing the signal to noise ratio for a given dose rate. To capture dynamics at high speed, the camera frame rate should be high; however, this requires a trade-off to be made - as framerate increases, a higher dose rate is needed to maintain a minimum number of electrons per camera frame. For this reason, the camera framerate should be set to the lowest framerate that still captures the dynamic event of interest.
Capturing data from the exact onset of dendrite growth was made possible by the use of the LookBack™ feature of the K3® IS camera. This feature enabled data to start being saved to disk even after the initial dendrite growth had already been observed but allows for data capture through the completion of the reaction. Figure 1 shows 10 frames from a 1 s long video acquisition of dendrite growth where data was collected at 75 fps. The corresponding video of the data set is shown below Figure 1.
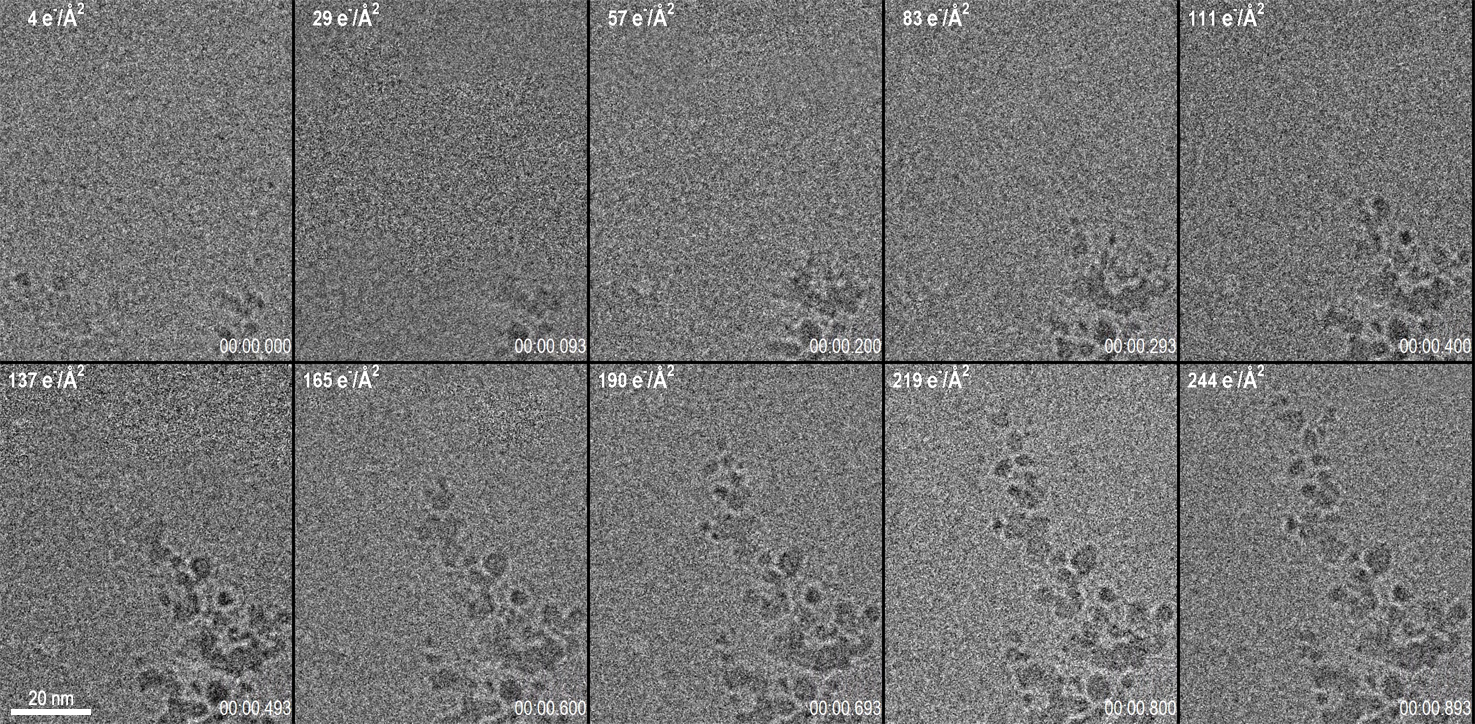
While the K3 IS can acquire data at frame rates in excess of 3500 fps, to set the optimal conditions for this experiment, a frame rate of 75 fps was chosen with a full 5760 x 4092 pix field of view. The dose rate of 270 e-/Å2/s was set at the sample, and the magnification was set to provide 0.63 Å pixels. This condition results in a very low dose rate at the camera of 3.6 e-/Å2/frame (e.g., 1.4 e-/pix/frame). For data visualization and analysis, the images were binned x2 to increase the signal to noise ratio at the expense of a small decrease in the spatial resolution. With the binned images, the dose rate per binned pixel was 5.7 e-/pix/frame. Given the Poisson statistics of the electron beam itself, this results in a signal-to-noise ratio for a single binned pixel of 2.3. Despite this very low signal-to-noise ratio of the incoming electron beam, the real-time, electron counting algorithm of the K3 IS enables accurate detection of this faint signal in the final image. By performing an exponentially weighted running average on the data to combine the signal from several frames, a clear video emerges. These calculations illuminate an important point for low-dose TEM: at low dose rates, 75 fps is nearing the maximum useful framerate from which information can be extracted at full resolution from every frame. Even a perfect detector, which determined the exact position of every electron striking it, would be limited by the incoming signal-to-noise of the electron beam itself as the framerate is increased.
The K3 IS uses a direct detection sensor and a real-time counting algorithm to come closer than ever before. The direct detection sensor eliminates sources of blurring and signal attenuation present in scintillator-based cameras. Meanwhile, the real-time counting algorithm significantly reduces the effects of noise sources within the camera, so that the primary noise component becomes shot noise from the electron beam itself. Unlike the K2® and K2 IS, the K3 IS brings the benefits of counting to in-situ video acquisition. The result is that low-dose videos like the ones seen here are made possible for the first time.
This result demonstrates that there are three basic parameters that must be properly balanced to provide adequate signal-to-noise in the incident signal from the electron beam: spatial resolution, temporal resolution, and electron dose rate (or flux). To maintain a useful signal-to-noise while pushing the temporal resolution higher, either the spatial resolution must be reduced, or the electron dose rate must be increased. In this experiment, a trade-off was made between the spatial resolution and the dose rate where the chosen condition collected data at 0.63 Å/pix and a dose rate of 270 e-/Å2/s. The next dataset also further reduces the dose rate to just 1.0 e-/Å2/s but has to do so with a slightly worse spatial resolution. However, this lower dose condition demonstrates another strength of the K3 IS: its large field of view and pixel resolution.
Observing a large field of view
Figure 2 shows data from a related, counted in-situ data set, also capturing the growth of a Cu dendrite. Again, the growth is beam induced. In this experiment, because the beam dose rate is lowered to 1.0 e-/A2/s, a slower growth rate of 10 nm/s is observed. For comparison, the frames shown in Figure 1 were captured in just 1 s at 75 fps, the data in Figure 2 was captured over 8 min (a duration almost 500 times as long). As a consequence of the longer experiment duration, the dose rate is 2 orders of magnitude lower in Figure 2 than in Figure 1, and 3 orders of magnitude lower than a single typical high-resolution TEM image. The total dose throughout the experiment is 474 e-/A2, or barely 2x that in the first video, and this dose provides the needed electrons to grow the dendrites to this size.
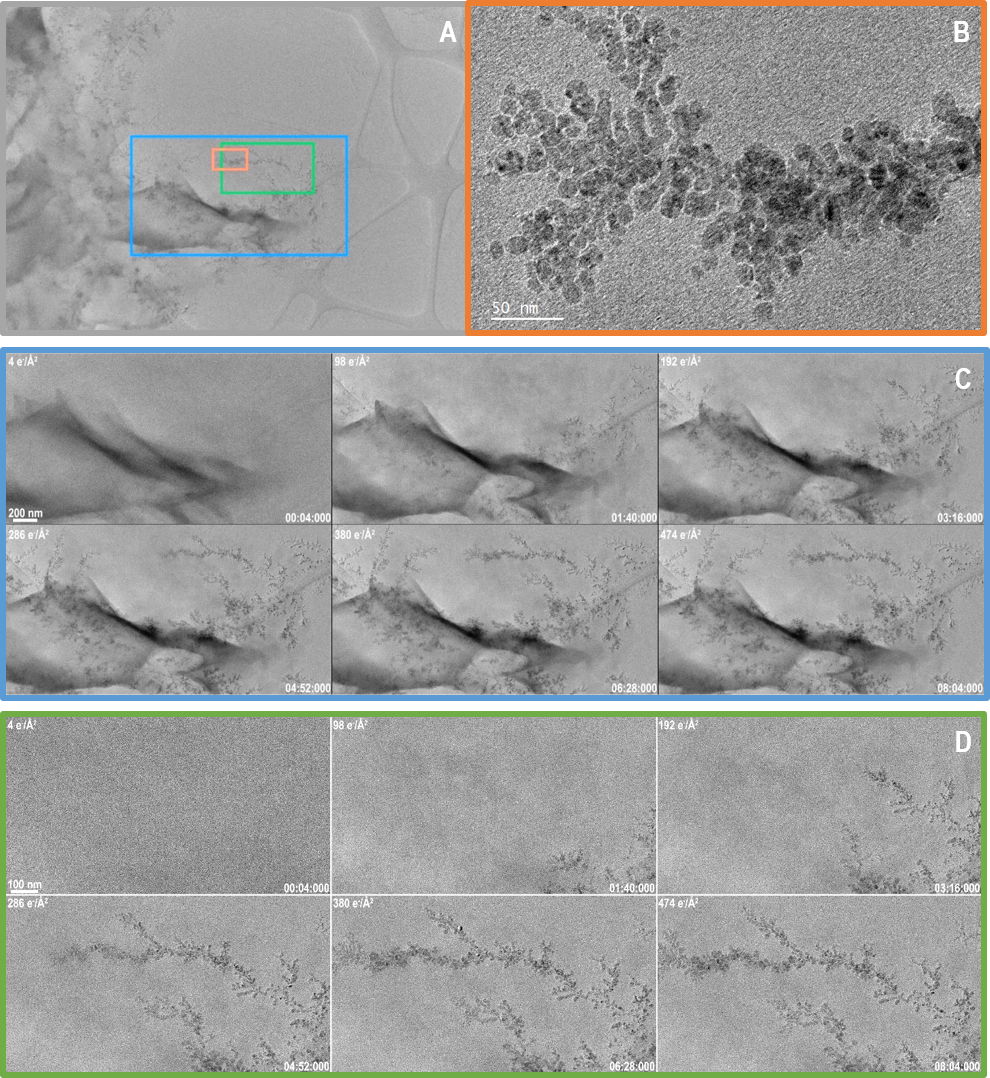

In addition to capturing data with a lower dose rate, the data shown in Figure 2 captured a much larger field of view of 5720 x 4060 nm or about 5.7 x 4.1 µm. This vast area is sampled with pixels that are just 0.5 nm across. For experiments like this one, where it is never certain a-priori where dendrites grow, there is great value in being able to acquire over a large area, but then process and analyze data from the locations where growth does occur. It is also possible to produce better statistics on the dynamic process since a large number of dendrites are simultaneously observed.
The counting algorithm employed by the K3 IS camera is ideal for this type of large field-of-view observation. For each raw camera frame, the location of individual incoming electrons is determined to sub-pixel accuracy. Although the K3 IS has >23.5 million pixels, this super-resolution mode results in images with >94.2 million pixels where this large field of view can be captured continuously at up to 30 fps.
Given the experimental trade-offs between electron dose rate, spatial resolution, and temporal resolution, the conditions for this experiment were to use a frame rate of 5 fps, an extremely low dose rate of 1.0 e-/A2/s, and pixel size of 0.5 nm. This combination yields a dose rate of 4.8 e-/pix/frame, slightly lower than in the high-speed data, where the dose rate per binned pixel per frame was 5.7.
Data analysis
The data presented in Figures 1 and 2 were processed using drift correction, binning, and an exponentially weighted moving average (over time, not space). However, the next step toward reaching a scientific conclusion based on the data was to perform an analysis of dendrite growth rates.
The DigitalMicrograph software has many tools for processing and visualization available from the standard user interface but has few tools for analysis. To supplement the user interface with additional flexibility and power, DigitalMicrograph has always had scripting capabilities (typically called DM-scripts). One great advantage of scripting within DigitalMicrograph is that the results of scripted acquisition or processing can then be visualized and manipulated using the standard user interface. However, the scripting language has been DigitalMicrograph-specific. While this means that it is uniquely tailored to the needs of microscopists, it also has limited the use of other open-source libraries within DigitalMicrograph during TEM data collection. Starting in DigitalMicrograph version 3.4.0, Gatan has integrated Python scripting the DigitalMicrograph adding the capability to edit and run Python (v 3.7) code directly from the scripting interface.
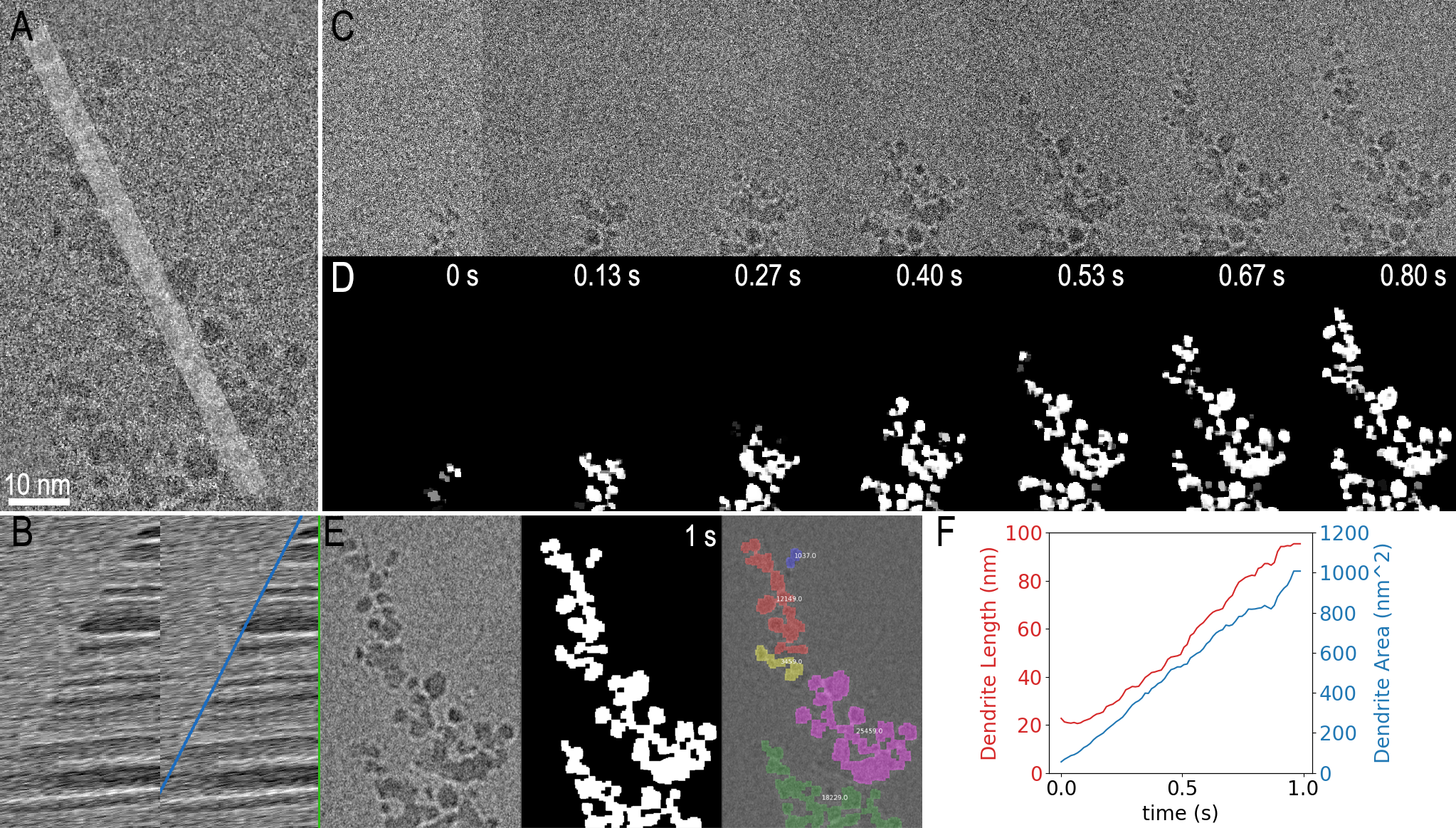
In Figure 3, two examples are shown using Python within DigitalMicrograph to analyze the dendrite growth data from Figure 1. In the first example, the code simply extracts a one-dimensional profile (shown in 3A) from the image as a function of time and presents this as a kymograph. The resulting kymograph is shown in part F, where a clear boundary, with its slope representing the average growth rate, can be observed. The profiles were measured live as the data was played back. It could also have been measured during data acquisition.
In the second analysis example, the same dendrite growth is measured using a more sophisticated image segmentation technique implemented with thresholding and morphological image processing functions from the scikit-image Python package. The input data (Figure 3 A and C) was noisy, so it was pre-processed within the same script. The final segmented result for selected frames is shown in Figure 3D, and 3E additionally gives a colored overlay on top of the filtered last frame. Various measurements can then be easily obtained from the segmented frames using another function in the scikit-image package. Figure 3 F shows 2 of the resulting measurements where both the length and area of the dendrite are plotted over time. On a PC with an Intel Core i7 8850H at 2.6 GHz, this entire process took <10 s, or 0.4 s per million image pixels. While this is far too slow to process all the data coming from the K3 IS camera in real-time, a subset of the data could be processed live in this way.
Conclusions
Beam-induced growth of Cu dendrites was captured at two very different time scales, both using the K3 IS in counted in-situ mode. In both cases, the incoming signal reaching the camera was very weak. At high speed, this was due to trade-offs necessary to balance spatial resolution, framerate, and electron dose rate. At slower speeds, the weak incoming signal was primarily due to the very low dose rate of just 1 e-/A2/s, which is three orders of magnitude less than for a typical HRTEM image. While small, these dose rates were still large enough to drive dendrite growth, though the growth rate was much smaller for the lower dose rate. The integration of Python in DigitalMicrograph enables analysis of this growth, and in the future, such analysis can be performed on a subset of the raw data in real-time during acquisition.
Acknowledgments
This work made use of the EPIC facility of Northwestern University’s NUANCE Center, which has received support from the Soft and Hybrid Nanotechnology Experimental (SHyNE) Resource (NSF ECCS-1542205); the MRSEC program (NSF DMR-1720139) at the Materials Research Center; the International Institute for Nanotechnology (IIN); the Keck Foundation; and the State of Illinois, through the IIN
The sample used in this work was provided by Kun He (NUANCE).
Data was collected by Ben Miller (Gatan), Paul Smeets, and Roberto dos Reis (NUANCE).