High-speed 4D STEM diffraction analysis of directionally-grown ZnO nanowires: benefits of a high-speed camera
Gatan instrument used
OneView® IS is the fastest and highest performance fiber coupled CMOS camera that captures 16 MP images at up to 25 fps. Using the STEMx™ system, the speed of the scanning probe is synchronized to the frame rate of the camera to collect 4D STEM diffraction datasets at full speed (up to 300 fps at 512 x 512 pixels).
Background
One of the main limiting factors in the conventional 4D STEM technology is the speed at which diffraction images can be acquired at each pixel in the STEM image. This speed is mainly limited by the frame rate of the conventional CCD cameras (30 fps for 512 x 512 images) and the speed by which 4D STEM datasets can be written and saved on disk. Such slow speeds limit the amount of data researchers can collect in a given amount of time, and are not feasible for beam sensitive specimens with limited allowable dose before they are damaged or structurally transformed.
Materials and Methods
Cross-section specimens isolating individual ZnO nanowires (NW) were selected for these experiments. As shown in a previous publication [Adv. Mater. Interfaces 2016, 1500598], the growth direction of these nanowires, on a GaN substrate, can be guided as a function of substrate Ga-ion bombardment, which results in different local surface boundaries to partition NW heterojunctions into different segments and orientations. Previously, HRTEM measurements determined tolerance of crawling nanowires to strain and lattice mismatch. However, conclusive quantitative strain measurements have proven difficult due to the beam-sensitive nature of the specimen and lack of a statistically sufficient number of diffraction measurements from the substrate-NW interface region. Using STEMx system with the OneView IS camera, two (256 x 85 x 1024 x 1024) datasets were collected each in <2 min, from NWs grown on bombarded and non-bombarded substrates. In addition to the visual comparison of averaged diffraction patterns at different interface regions, 2D strain maps were calculated along orthogonal axes within the diffraction pattern and with reference to the unstrained GaN substrate. Using the STEMx Analytical Suite, the relative in-plane, out-of-plane, shear and rotation at the interface can be quickly gauged for large datasets (each >40 GB). Overall, the current strain mapping results show the character and degree of lateral strain (ε_uu) more relaxed in directions parallel to the nanowire long axis through the ion bombarded regions (Figure 1) compared to the localized intense lateral strain points perpendicular to the same axis (ε_uu in Figure 2). Alternatively, the relative degree of compressive strain (ε_vv) was observed to be about the same in either case.
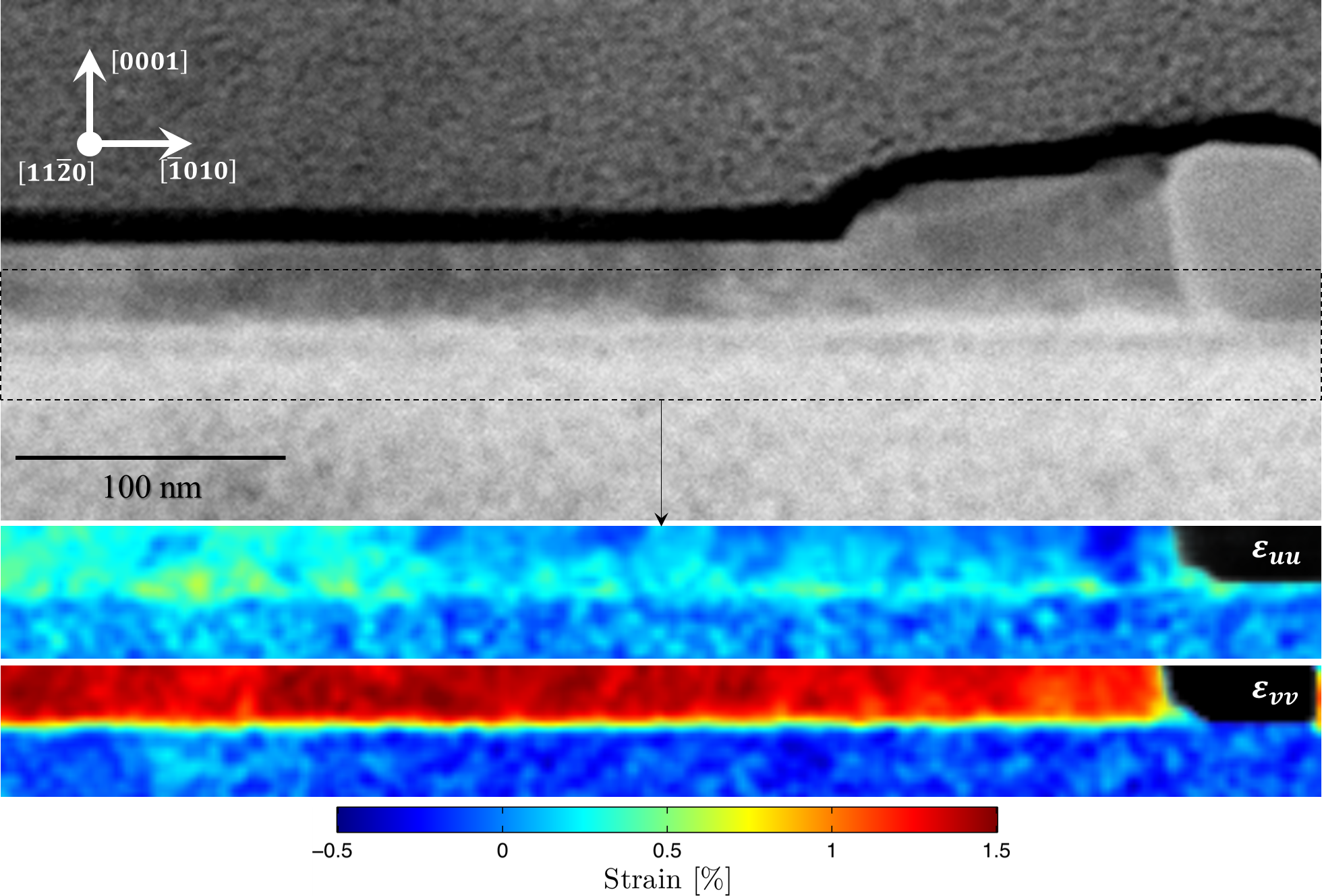
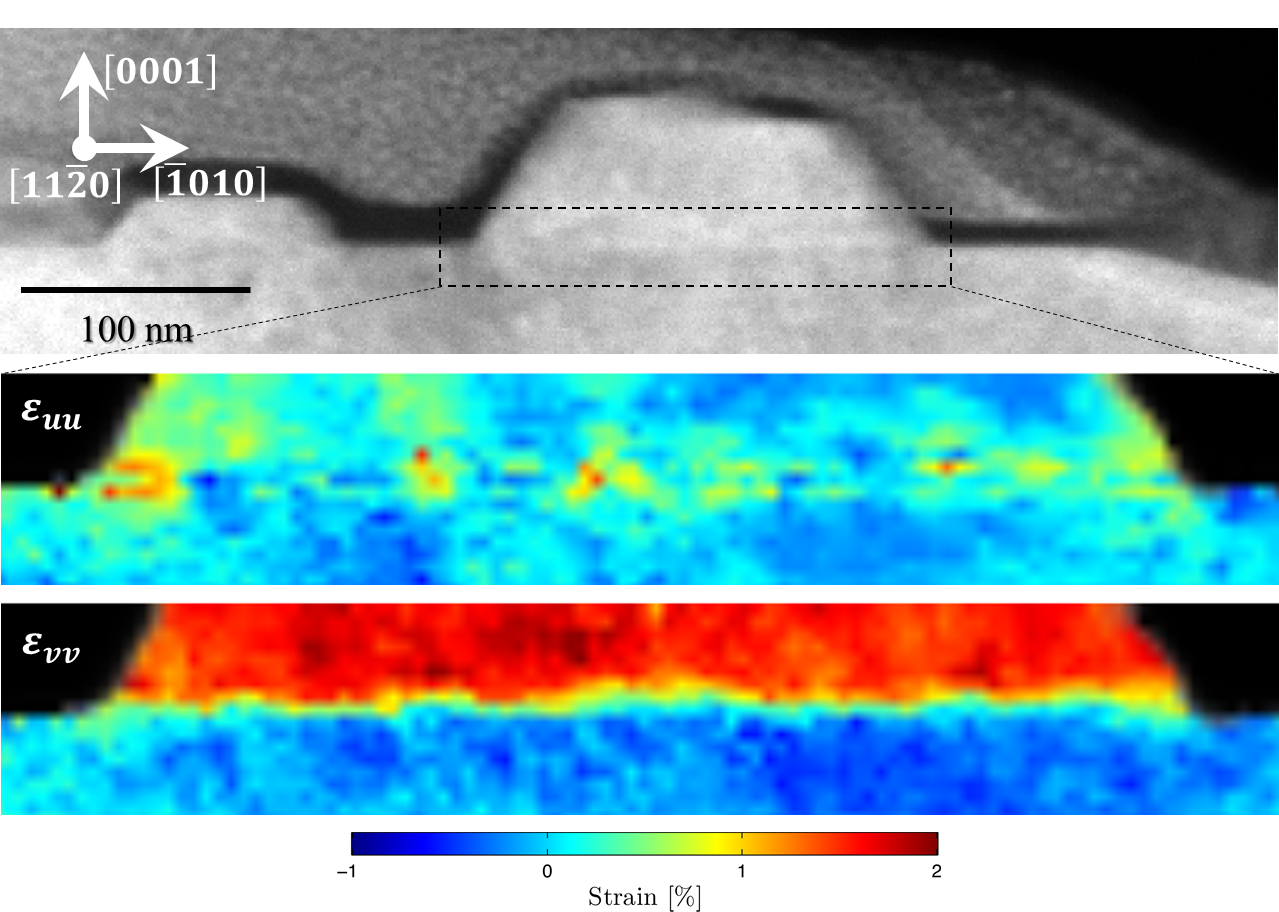

Further data collection is needed to further support these conclusions. Current experiments hint at new avenues to explore in understanding and controlling lateral epitaxy of nanocrystals. The implications of these results can be wide-ranging; touching areas of future fabrication of nanowire-based devices. For instance, in combining top-down and bottom-up fabrication techniques or incorporating II-V and III-nitride semiconductors into industrial processes.
Summary
By using the STEMx system, the fast frame rate OneView camera was applied to drive the STEM probe on beam sensitive ZnO NWs, grown on GaN substrates. High-speed 4D STEM data acquisition significantly reduced the specimen damage previously encountered during data collection (saving time and money). The data acquired with the OneView camera provides high-resolution information capable of direct observation of single systems over large length-scales. Through intelligent modeling of this data, it also enables a visualization of nanoscale effects in an overall microsystem. Such results produce insights into the mechanisms of nano epitaxial growth and foundational information to incorporating these systems into future technologies.
Credit(s)
Special thanks to Dr. E. Garratt (NIST), Dr. A. Herzing (NIST), Dr. C. Ophus (NCEM) and Dr. J. Ciston (NCEM).